|
Longitudinal studies in wild animals are thus important to precisely assess prevalence of infected animals and its temporal variation. Important seasonal variation in the prevalence of infected animals can depend on the period the samples were collected, and, in turn, can lead to misrepresentative conclusions regarding the level of bat exposure to viruses, in particular in cross-sectional studies. For example, seasonality has been identified as a major driver of the infection dynamics of many pathogens, affecting both host susceptibility and transmission. Differences between locations, sex, and bat species may be explained by a range of factors. Interestingly, higher detection rates were found for both viruses in bats using caves as day-roost sites, suggesting that cave-roosting behavior maybe favorable for horizontal transmission between bats. The overall proportion of positive bats detected for either AstVs or PMVs was consistent with previous studies performed in the WIO region, and in other tropical regions. The highest AstV detection rate in Mozambique was observed in Triaenops afer (68.6% ± 12.7%). On Madagascar, a high detection rate was found in species of the genus Miniopterus as compared to the other taxa (χ 2 = 162.4, df = 1, P < 0.05), and especially in Miniopterus manavi (88.9% ± 14.5%), Miniopterus gleni (86.7% ± 17.2%), and Miniopterus sororculus (71.4% ± 33.5%). AstV prevalence was significantly higher in bats species roosting in caves (27.5% ± 3.9%) than in buildings (2.5% ± 2.0%). Positive samples were detected only in Mozambique (20.2% ± 3.6%) and on Madagascar (18.6% ± 3.5%), without significant variation between these two locations (χ 2 = 0.3, df = 1, P > 0.5). One hundred and forty-two samples tested positive for AstV (mean detection rate ± 95% confidence interval: 16.3% ± 2.5%) (Additional file 2). Phylogenetic trees were generated by maximum-likelihood using PhyML software 3.1, with a GTR evolutionary model, and 1000 bootstrap replicates. 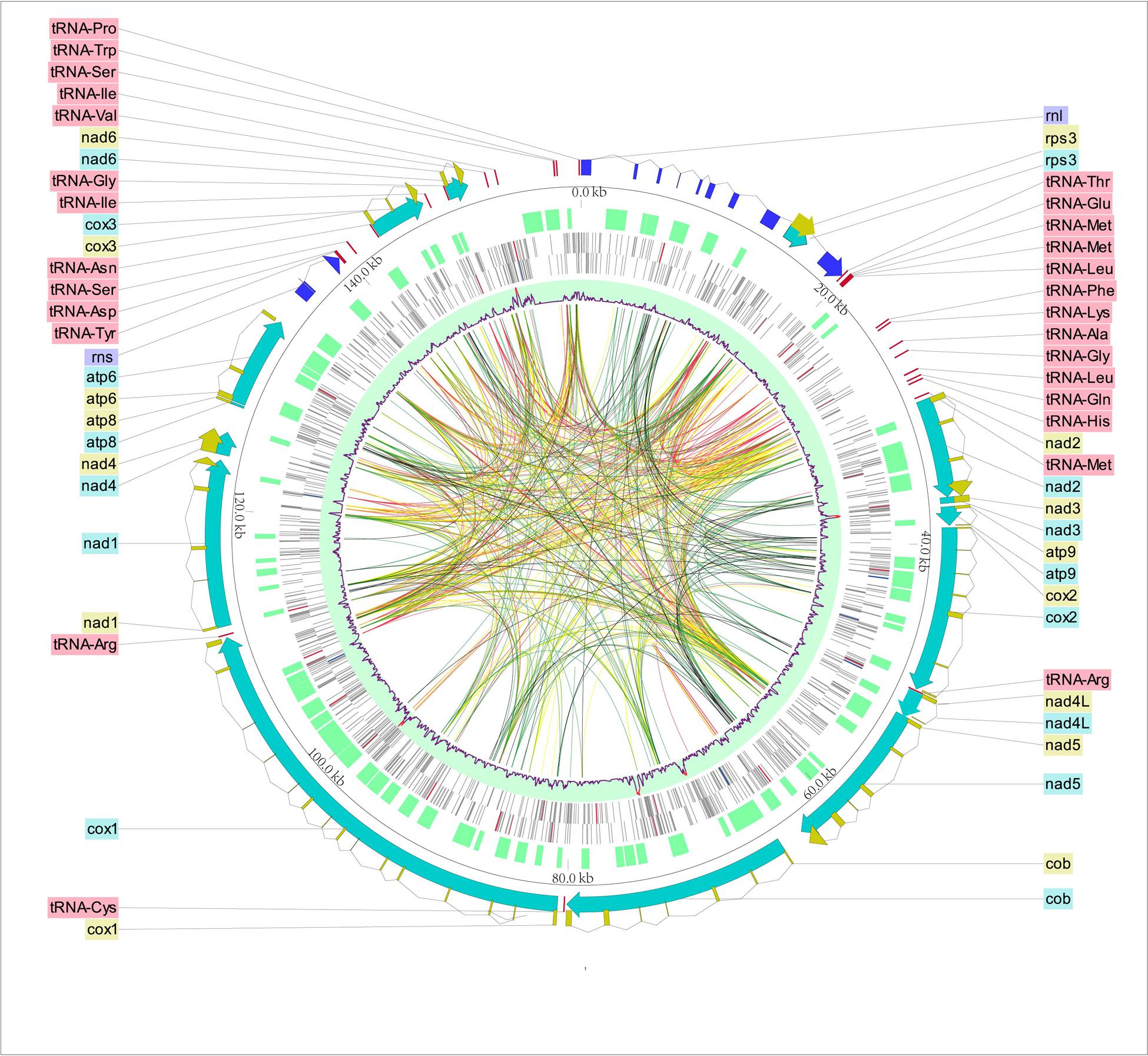
Then, AstV and PMV sequences generated in this study were respectively aligned with 105 and 74 reference partial nucleotide sequences, using CLC Sequence Viewer version 7.6.1 (CLC Bio, Aarhus, Denmark). Sequences were compared to reference sequences in NCBI GenBank using the Basic Local Alignment Search Tool (BLAST) with the standard nucleotide BLAST (BLASTn) algorithm (BLAST was performed on August 18th, 2021). Genetic diversity was explored with pairwise distance values obtained from phangorn package in R, version 2.6.3. The 33 partial AstV sequences and 13 partial PMV sequences generated in this study were deposited in GenBank respectively under the accession numbers MZ614404 to MZ614436 and MZ614437 to MZ614449. Nucleotide sequences were aligned to generate consensus sequences, and were edited manually using ChromasLite 2.6.5 (Technelysium Pty, South Brisbane, Australia). PCR products of expected size were submitted for direct Sanger sequencing (Genoscreen, Lille, France). Analyses were conducted with R, version 4.0.5. country or island), on virus detection, and to investigate potential associations between AstV, CoV, and PMV. cave, building, tree), host species, sex, and sampling location (i.e. Pearson Chi square tests were conducted to examine the effect of the roost sites (i.e. PCR products were visualized on 2% agarose gels stained with 2% Gelred (Biotium, Hayward, CA, USA). Molecular detection was performed using semi-nested polymerase chain reactions (PCRs), as previously described. Additional assays were thus performed for the detection of the AstV (355 samples) and PMV (704 samples) RNA-dependent RNA-polymerase (RdRp) genes. All samples were previously tested for the presence of CoV some of them were also tested for the presence of AstV (516 samples ) and PMV (167 samples ). bat species, location, date, type of samples) is provided in the Additional file 1. 
The list of samples included in this study (e.g. Biological material was collected on Madagascar, in Mozambique, on Mayotte, and on Reunion Island as part of previous investigations on infectious agents circulation in bats (details relating to the collection of biological material are available in ).
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
